Date: 16-24 October 2015
Venue: Nha Trang University, 02 Nguyen Dinh Chieu, Nha Trang, Vietnam
Instructors:: Dr. Binh Thuy Dang, Nha Trang University (contact to discuss content), Dr. Jonathan Puritz, Harte Research Institute, Texas A&M Corpus Christi, Dr. Chris Bird/Sharon Furiness, Harte Research Institute, Texas A&M Corpus Christi, MSc. Vu Dang Ha Quyen, Nha Trang University (contact to lab work)
Course coordinator. Binh Thuy Dang
Workshop description and aims:
This training- workshop will bring together trainers involved in teaching NGS analysis to life scientists and give them the opportunity to work on developing a unified collection of training materials, tailored to different audiences and updated collaboratively.
Course Description
This intensive 8 days course will introduce attendees with a strong biology background to the practice of analyzing short-read sequencing data from Illumina and other next-gen platforms. The first day will introduce students to basic concept of NGS, computational thinking and large-scale data analysis on UNIX platforms. The following 2 days will practice Ezrad technique in the Molecular lab. The last 5 days focus on assembly, and analysis of short-read data for SNPs calling and population structure.
Participants:
Members of the project "Peer Cycle 3" from Nha Trang University (NTU), Can Tho University (CTU), Inland Fisheries Research and Development Institution (IFReDI), Living Aquatic Resource Research Center (LAReC), Others if possible The expected number is 16-20.
Learning outcome
By the end of the course, students will be able to
- Understand some principles behind NGS, and know the challenges created by NGS
- Conduct EZRAD technique; DNA extraction, DNA qualification, libraries preparation
- Understand the uses of NGS data in: filter data, create reference, SNPs calling and analysis, population structure analysis.
- Know how to use the software commonly used in bioinformatic
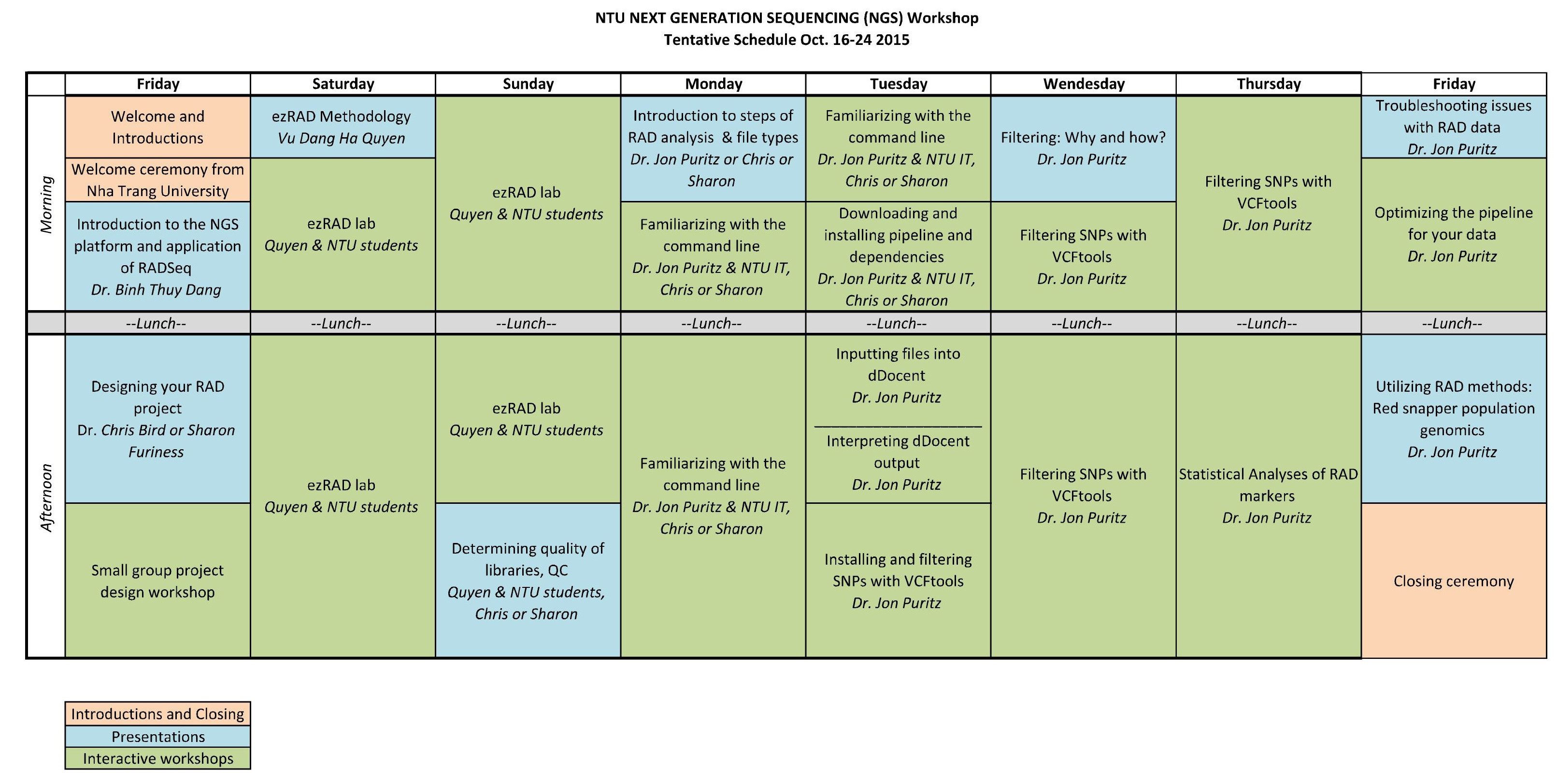
Workshop Schedule